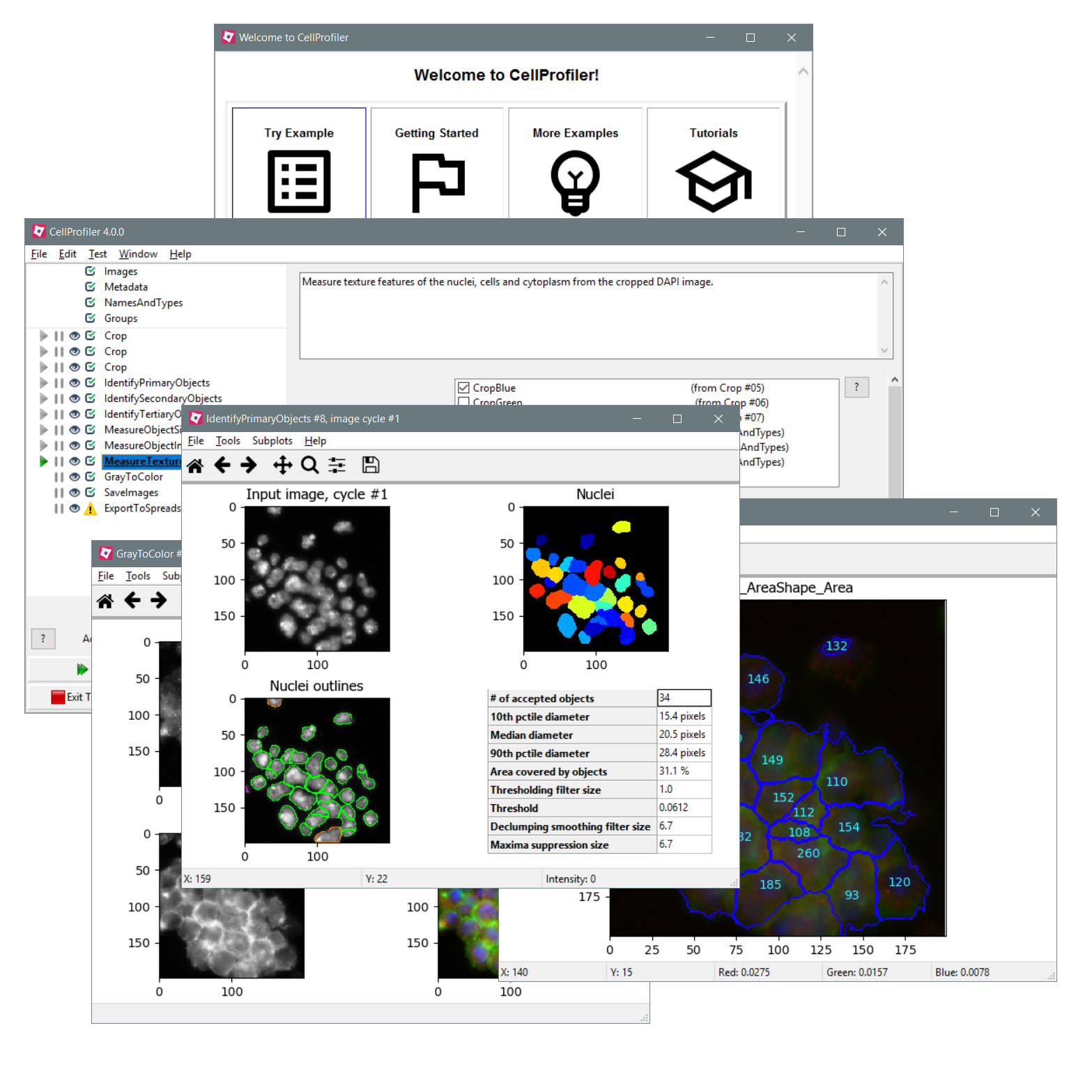
As we continue to make changes and streamline, we’re going to use the data from 3.0’s new telemetry feature to see what modules and settings people generally use in pipelines and which seem to be superfluous we really therefore appreciate if you turn it on (either on startup or in the preferences menu) so we can see (and not remove!) what’s most important to you in the community.ġ) These are the major pipeline-breaking changes to keep an eye out for:Ĭhanges to how thresholds are calculated*- these will affect the Threshold module as well as IdentifyPrimaryObjects and IdentifySecondaryObjects.ĬellProfiler 2.X’s way of calculating a 2-class Otsu threshold turned out to be non-standard we’ve therefore started using the implementation in scikit-image instead to bring it in line with how other image processing tools calculate it. If you find something you’ve used before is now broken and you need help fixing it, please do make a new forum thread and let us know and we’ll do everything we can to help you find a new solution. We’ve highlighted the biggest changes at the top for your convenience. You probably will have to spend a bit of time optimizing and re-checking pipelines when you DO make the switch, but we truly believe you’ll find it well worth your while in the long run. We’ve therefore pulled together a list of most of the major and minor changes you’ll encounter when switching between 2.2 and 3.0. In order to facilitate the speedup and continue the process of streamlining the code, a few things had to go we also removed some things we felt were causing “option fatigue” for the sake of user friendliness going forward. Threshold:[module_num:15|svn_version:'Unknown'|variable_revision_number:12|show_window:True|notes:['The Watershed module finds objects that have bright signal, so the cytoplasm that will define the cell volume should have bright signal.For those of you who’ve been with us for a long time though, the obvious next question after how to use the new test mode is will my old CellProfiler pipelines work in the new version? We feel the same way – the pipelines you’ve accumulated over the years are precious resources! The good and bad news is that the answer is Yes, mostly. Select the input objects:ErodedResizedNucleiObjects Name the output object:ErodedResizedNucleiObjectsĬonvertObjectsToImage:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] ResizeObjects:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False]ĮrodeObjects:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] Watershed:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] RemoveHoles:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] Two-class or three-class thresholding?:Two classesĪssign pixels in the middle intensity class to the foreground or the background?:Foreground Select the measurement to threshold with:None Lower and upper bounds on threshold:0.0,1.0 Threshold:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] MedianFilter:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] Select the image with the desired dimensions:None Resizing method:Resize by a fraction or multiple of the original size Resize:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] Select image to match in maximum intensity:None 
Intensity range for the output image:0.0,1.0 Intensity range for the input image:0.0,1.0 Upper intensity limit for the input image:1.0 Lower intensity limit for the input image:0.0 Method to calculate the maximum intensity:Custom Method to calculate the minimum intensity:Custom Rescaling method:Stretch each image to use the full intensity range RescaleIntensity:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] Groups:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] Select the rule criteria:and (metadata does ChannelNumber "0") Select the rule criteria:and (metadata does ChannelNumber "1") Select the rule criteria:and (metadata does ChannelNumber "2") NamesAndTypes:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] Select the filtering criteria:and (file does contain "") 
Regular expression to extract from folder name:(?P)$ Regular expression to extract from file name:^(?P.*)_xy(?P)_ch(?P) Metadata extraction method:Extract from file/folder names Metadata:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False] Select the rule criteria:and (extension does isimage) (directory doesnot containregexp "\\\\.") Images:|batch_state:array(, dtype=uint8)|enabled:True|wants_pause:False]
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
